Behavior Atlas:
Get started now GitHub repository
Introdution
Behavior Atlas is a spatio-temporal decomposition framework for automatically detecting behavioral phenotypes. It receives 3D/2D continuous multidimensional motion features data input, and unsupervisedly decompose it into understandable behavioral modules/movements (e.g., walking, running, rearing). Our framework emphasizes the extraction of the temporal dynamics of movements. Without making model assumptions, similar movements with various time durations and temporal variability can be efficiently detected. Besides the decomposition, the constructed self-similarity matrix of these movement segments describes the structure of involved movements. Further dimensionality reduction and visualization enable us construct the feature space of behavior. This helps us study the evolution of movement sequences for higher-order behavior and behavioral state transition caused by neural activity.
Features
1. The real 3D reconstruction of animal poses with a high signal-to-noise ratio
- a. Capturing Image streams from four cameras with different 2D views;
- b. Using DeepLabCut (DLC) to track animal body parts and generate 2D skeletal trajectories separately (color-coded traces);
- c. Reconstructing 3D body skeleton by integrating these four data streams (based on pose3d).
| Video1 | 3D reconstruction demo video, Download this video Video1-3D_Capture_and_recons_demo.mp4 (6.91MB) |
| Figure1 | The pipeline of 3D animal skeletal reconstruction. |
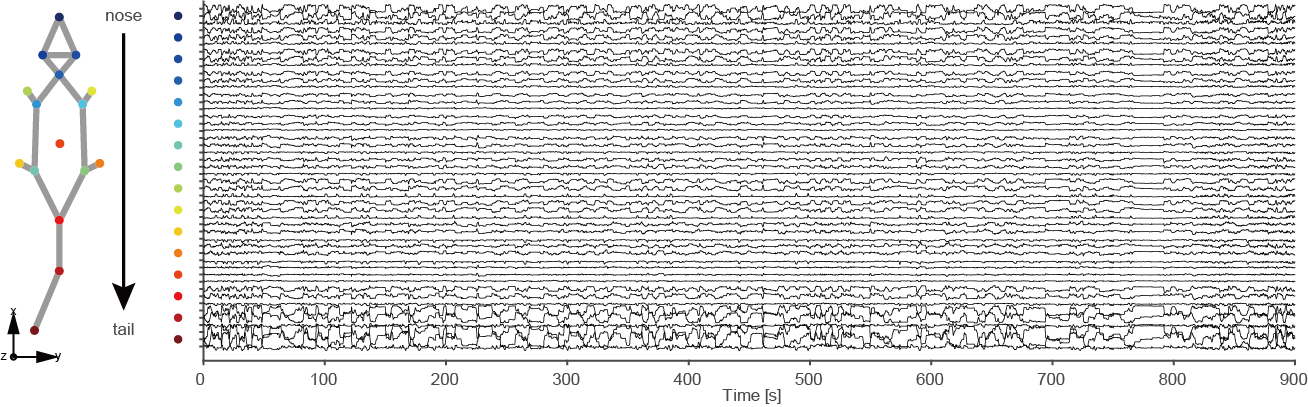
| Figure2 | Representative (900s) mouse body tracking trace data showing with 48 data vectors obtained by 3D reconstruction. |
2. Parallel and two-layer framework for motion feature decomposition.
There are many similar previous works such as MotionMapper, Moseq were used to detect stereotyped movement modules on flies or mice. Our framework focus on address the high-dimensionality and large temporal variability of rodent behavior. we adopted a parallel motion decomposition strategy following the natural characteristics of animal motion. That is, two-layer dynamic temporal decomposition is performed on the continuous animal skeleton postural data to obtain Non-locomotor movement (NM) space. Finally, together with locomotion dimension, unsupervised clustering is used to reveal the structure of behavior.
| Figure3 | Spatio-temporal decomposition framework for animal behavior analysis. |
3. Mapping Mouse Movements with Low-Dimensional Embeddings and Unsupervised Clustering
We demonstrate our framework in 15-minute experimental data collected in a featureless circular open-filed. Thus, we obtained 11 types (determined by Bayesian Information Criterion ) of movement.
| Video2 | Decomposition demo of 15-minute experimental data collected in a featureless circular open-filed Video2-decomposition_demo.mp4 (8.92MB) (CQI: Clustering Quality Index helps us determine the stereotyped/non-stereotyped movements. CQI is evaluated by integrated the intra-cluster and inter-cluster correlation coefficients. Larger CQI indicates more stereotyped movement segment) |
We examined the video clips in each group, post-hoc manually labeled annotated as running, trotting, stepping, sniffing, rising, right turning, up stretching, falling, left turning, walking.
| Up stretching | Running | Rising | Sniffing |
|---|---|---|---|
| Stepping | Falling | Diving | Right turning |
| Trotting | Left turning | Walking | |
The 15-minute experimental data were decomposed into 936 NM bouts, and embedded these bouts in a low-D space:
- a. Firstly, a 936×936 segment kernel matrix was constructed using the DTAK metric. This segment kernel matrix can flexibly represent the relationship and provide valuable insights between each behavioral component sequence in their feature space.
- b. Because of the high-dimensionality of behavior, we adopted an algorithm called UMAP which can preserve both local and global structure of the dataset. Provide us a 2D embeddings of these NM segments.
- c. The combination of 2D NM space and motion dimension provide a 3D movements space representation.
| Video3 | Spatio-temporal feature space of behavioral components Video3-cluster_embeddings.mp4 (7.61MB) |
About the project
License
Behavior Atlas is distributed by an MIT license.